Hidden Viral Signals in SARS-CoV-2 Reveal New Ways the Virus Evades Our Immune System
Article audio
Listen to a generated narration of this post.
When we think about how SARS-CoV-2 evolves and adapts, most attention goes to changes in the spike protein—the viral key that unlocks our cells. But scientists have now discovered a hidden viral signal within the virus’s nucleocapsid gene that produces a smaller, truncated protein. This tiny viral trick helps the virus dodge our immune defenses in ways we hadn’t fully appreciated before, offering fresh insights into how COVID-19 variants might gain an edge.
TL;DR
- A new transcription-regulating sequence inside the SARS-CoV-2 nucleocapsid gene creates a truncated nucleocapsid protein (N*) that boosts viral fitness.
- The N* protein interferes with host antiviral defenses, including interferon responses and ribonucleoprotein granule formation, helping the virus evade immune detection.
Coronaviruses like SARS-CoV-2 have relatively large RNA genomes, but they pack a lot of information into a compact space. The virus relies on special RNA signals called transcription-regulating sequences (TRSs) to produce subgenomic mRNAs, which in turn make the proteins needed for replication and interaction with host cells. While much research has focused on mutations in the spike protein, changes in other parts of the genome, such as the nucleocapsid (N) gene, can also influence how well the virus replicates and avoids immune responses. Until recently, these non-spike changes remained poorly understood.
Two recent studies published in PLOS Biology used a combination of genetic engineering and cell culture experiments to investigate a novel TRS that appeared within the SARS-CoV-2 nucleocapsid gene early in the pandemic. By creating recombinant viruses with and without this internal TRS—and by separating its effects from accompanying amino acid changes in the full-length N protein—researchers could pinpoint how the new signal influenced viral replication and immune evasion. They also examined how the truncated N* protein affected antiviral pathways, including interferon signaling and the formation of ribonucleoprotein granules that help cells respond to viral infection.
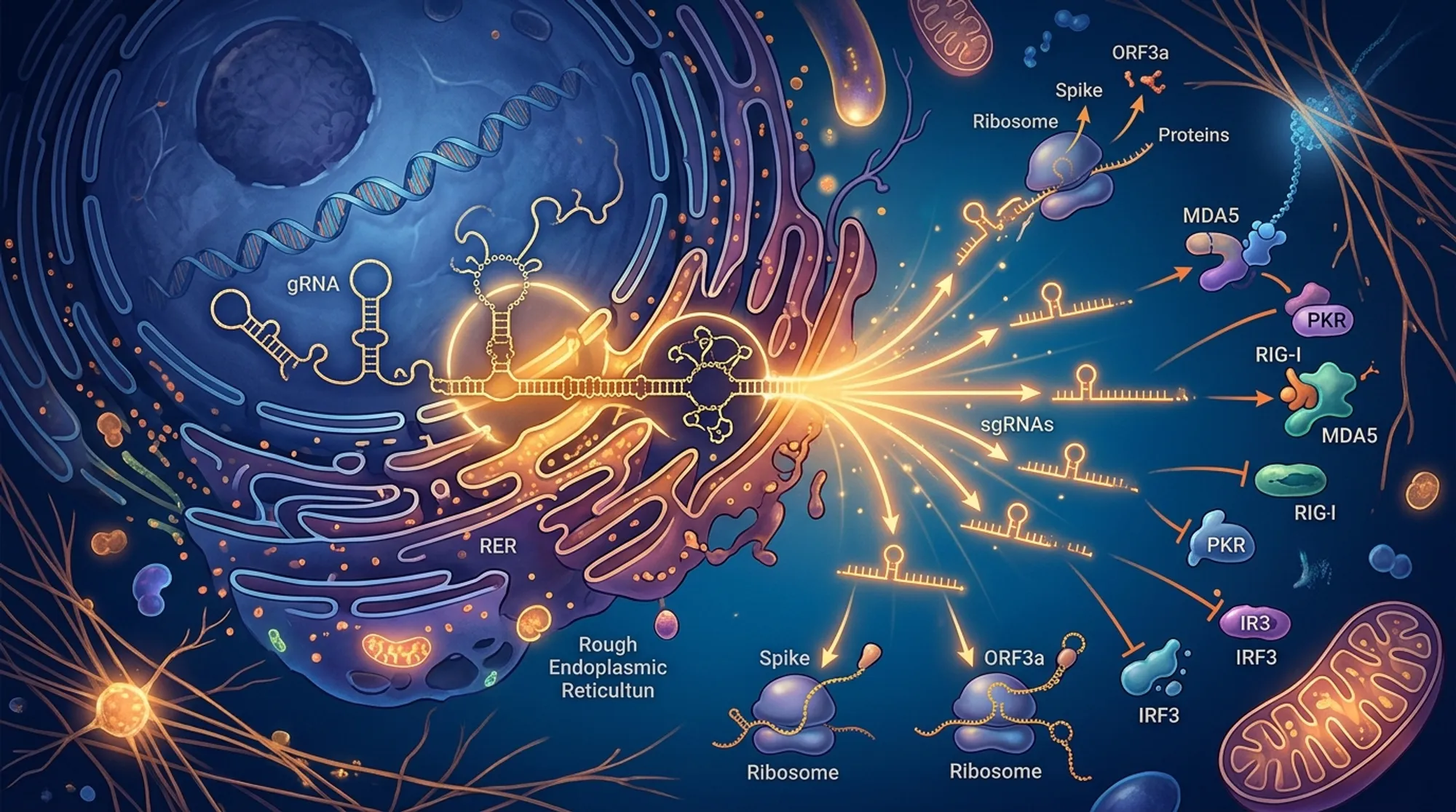
The studies showed that the internal TRS leads to production of a truncated nucleocapsid protein, called N*, which contains a double-stranded RNA binding motif. This N* protein binds viral RNA and blocks key antiviral responses. For example, it inhibits the activation of RIG-I, a sensor that detects viral RNA and triggers interferon production. Additionally, N* prevents the formation of stress granules and other ribonucleoprotein complexes that normally help the cell mount defenses against the virus. Viruses expressing N* outcompeted those without it in cell cultures, demonstrating a tangible fitness advantage. Interestingly, the presence of N* was found in multiple SARS-CoV-2 variants, including Alpha, Gamma, and Omicron, pointing to its evolutionary importance.
This discovery broadens our understanding of how SARS-CoV-2 adapts beyond the well-studied spike protein. The truncated N* protein represents a new viral tool that enhances fitness by dampening multiple host antiviral mechanisms. Recognizing the role of such non-spike mutations could inform future antiviral drug development and vaccine strategies that target other viral components. Moreover, these findings highlight the complexity of virus–host interactions and the sophisticated ways viruses evolve to survive within human populations.
While these findings are compelling, much of the evidence comes from cell culture models, and confirming the effects of the N* protein in living organisms remains an important next step. The exact composition and antiviral functions of ribonucleoprotein granules are still not fully understood, making it challenging to dissect all mechanisms by which N* enhances viral fitness. Additionally, the impact of N* may vary depending on other mutations present in circulating variants, underscoring the need for ongoing research to track how these viral adaptations influence disease outcomes.

